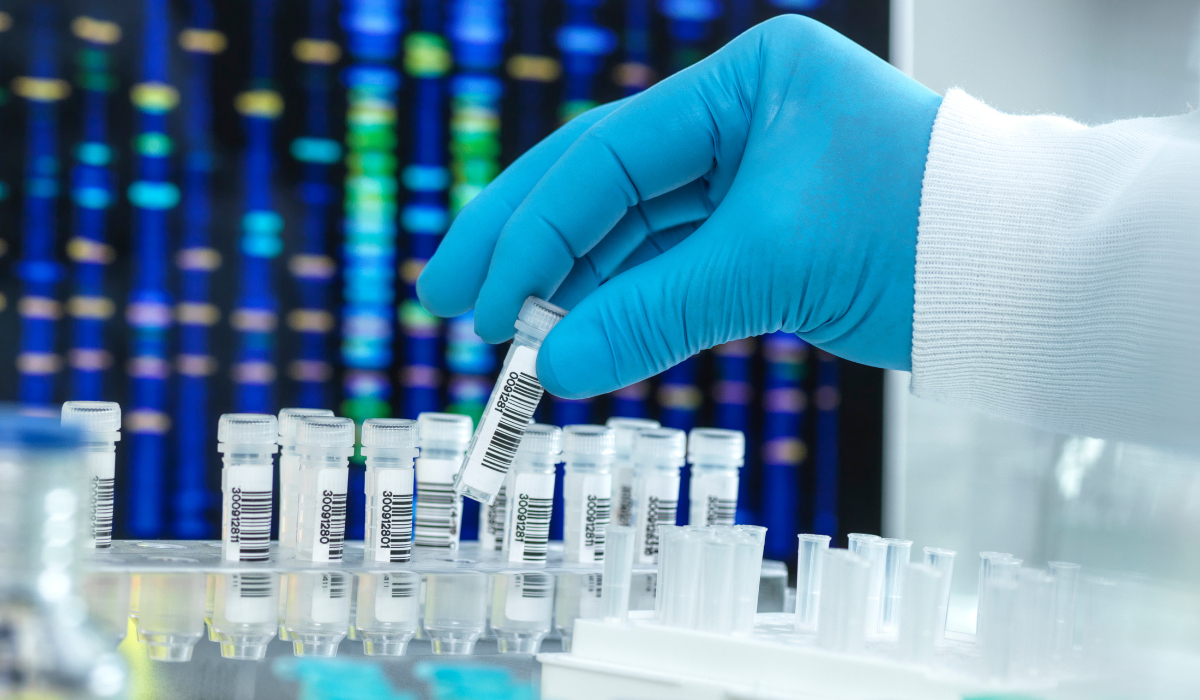
The field of genomics has expanded rapidly in the last two decades thanks to the advent of next-generation DNA sequencing (NGS) technologies. These technologies automate the sequencing and analysis of DNA, providing high throughput data acquisition. Illumina is the market leader in NGS, whose technology is based on sequencing-by-synthesis of randomly digested short fragments in nanostructured flow cells. Its latest sequencer, the NovaSeq X, can run 128 genomes per day, at an approximate cost of $200 per genome, after capital expenditures. Illumina devices excel at producing high quality short-read data (less than 1000 bases), with greater capacity for long-reads currently in development. Other molecular diagnostics technologies, such as the Sequel II from PacBio or the MinION and PromethION from Oxford Nanopore Technologies, specialise in much longer read-lengths, but with necessarily slower data acquisition times.
Transcriptomics
Transcriptomics is the study of the total set of expressed RNA in a cell, or population of cells, at a specific moment in time. Analysing a transcriptome provides a comprehensive understanding of which genes are active and producing RNA, as well as the quantity and variation of RNA molecules. This information helps in deciphering the underlying molecular mechanisms involved in various biological processes, such as development, disease progression and response to external stimuli. The mechanisms for the expression of DNA information into proteins are complex and highly regulated by processes not obvious from genomic information alone.
Transcriptomics is applied to the diagnosis and profiling of cancer through the identification of gene expression patterns associated with different types of cancer. It can also be applied to infectious diseases by analysing pathogenic transcriptomes or the host immune response and neurological disorders, by looking for expressed biomarkers in the brain or peripheral tissue, among others.
The technology already developed for DNA sequencing is similarly capable of sequencing RNA with modifications to the biochemistry, called RNA-seq. The major suppliers of NGS devices and services also provide kits for RNA-seq.
Proteomics
Proteomics is the study of the entire set of proteins produced by a cell or tissue at a single point in time. It involves the identification, characterisation and quantification of proteins, as well as the study of their functions and interactions within biological systems. The approach can also identify post-translational modifications that effect protein function.
Proteomics uses molecular diagnostic techniques common across biotechnology and life science laboratories, rather than relying on a handful of devices as is the case for genomics and transcriptomics. Mass spectrometry (MS) is the principal technique for identifying and characterising proteins. MS is not inherently quantitative, but labelling methods have been developed over the past decade to provide accurate quantification of protein expression. Quantitative proteomics is of particular importance when tracking disease progression and response to treatment over time.
There has been a recent push towards protein nanopore sequencing, analogous to DNA nanopore sequencing. However, polypeptides are at least an order of magnitude more complex than DNA and so present unresolved problems to sequencing. It is likely to be some time before the first desktop protein sequencing devices are commercially available.
Metabolomics
Metabolomics is the study of metabolites, small molecules within cells, tissues or biological fluids that are the products of cellular processes. It is the youngest of the ‘omics’ but shows the most promise for molecular diagnostics and personalised medicine in clinical practice. An understanding of the complete set of metabolites within a biological sample provides insights into the biochemical pathways that underlie cellular function. Metabolite levels and interactions change in response to genetic makeup, environmental influences or drug treatments and so can be excellent indicators of the presence of disease or the progression of treatment.
Metabolome biomarker discovery utilises common chemical analysis techniques, such as LC-MS and NMR, to characterise small molecule metabolites. Equipment of this kind requires large capital expenditure to purchase and significant expertise to operate, so metabolomic discovery is offered as services by research institutes or businesses, such as Biogenity and Metabolon.
The translation of metabolomics into clinical practice has been slow due to the differences in case-control studies used for discovery, versus the sample-control set testing used for diagnosis. Problems persist in the standardisation of methodologies between group studies and analytical approaches to gathered data, making accurate comparisons between different data sets difficult. This is especially problematic for personalised testing, which relies on robust data sets for comparison. Thus far, these concerns have prevented regulatory approval, with no FDA approved metabolomic tests currently available. This is, however, expected to change as the field matures and rigorous standards
are adopted.
The future transition from omics research into clinical diagnostics will focus on the identification of biomarkers that indicate the presence or progress of a disease. It is not economically viable to undertake a full suite of omics assessments for each patient, in most instances, so specific tests must be developed for specific diseases. Having considered the advances in omics over the last decade, the following will consider some of the future trends in molecular diagnostics technology.